(This work was done in collaboration with Dr. Robert Nowak.)
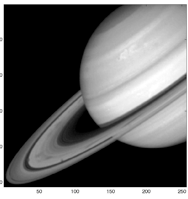
Much of my research has been focused on multiresolution methods for signal recovery from noisy observations, an important theoretical problem with a large number of diverse and critical applications. In the past decade, wavelet analysis and related methods have proved themselves to be good practical tools with a number of favorable theoretical properties, and many investigators have considered the use of wavelet representations for image denoising, deblurring, and tomographic image reconstruction. The effectiveness and theoretical optimality of these methods, however, are constrained by two common phenomena: (1) edges in images or lower-dimensional manifolds embedded in higher-dimensional observation spaces, and (2) the presence of non-Gaussian noise. Both are particularly prevalent in the context of photon-limited imaging, which arises routinely in applications such as bioimaging and astrophysics. The localization of singularity structures in multi-dimensional data is critical to many information processing tasks, including estimation, compression, and classification. Ignoring edges and singularities can severely hamper performance, but computationally tractable and accurate extraction of sub-manifolds is a challenging open problem. Similarly, ignoring the effects of non-Gaussian noise can be a significant source of error, but more precise noise models can lead to highly non-linear computational problems. Both model accuracy and computational efficiency must be considered in the development of effective, practical information processing tools. In collaboration with Prof. Robert Nowak, I demonstrated that fast and practical platelet methods effectively address both these problems, can nearly achieve theoretical lower bounds on the best possible error convergence rates, and perform very well in practice.
Platelets are localized basis functions at various locations, scales and orientations that can produce highly accurate, piecewise linear approximations to images consisting of smooth regions separated by smooth boundaries. Moreover, platelet representations are especially well-suited to the analysis of Poisson data, unlike most other multiscale image representations, and they can be rapidly computed. I have shown that that platelet-based estimators nearly achieve theoretical lower bounds on the minimax error convergence rate and can significantly outperform conventional wavelet and wedgelet approximations.
Download information:
This toolbox contains MATLAB files for platelet image reconstruction under Poisson or Gaussian noise models. To get started, try running Example.m. Thank you to Drew Miller for this new fast version!